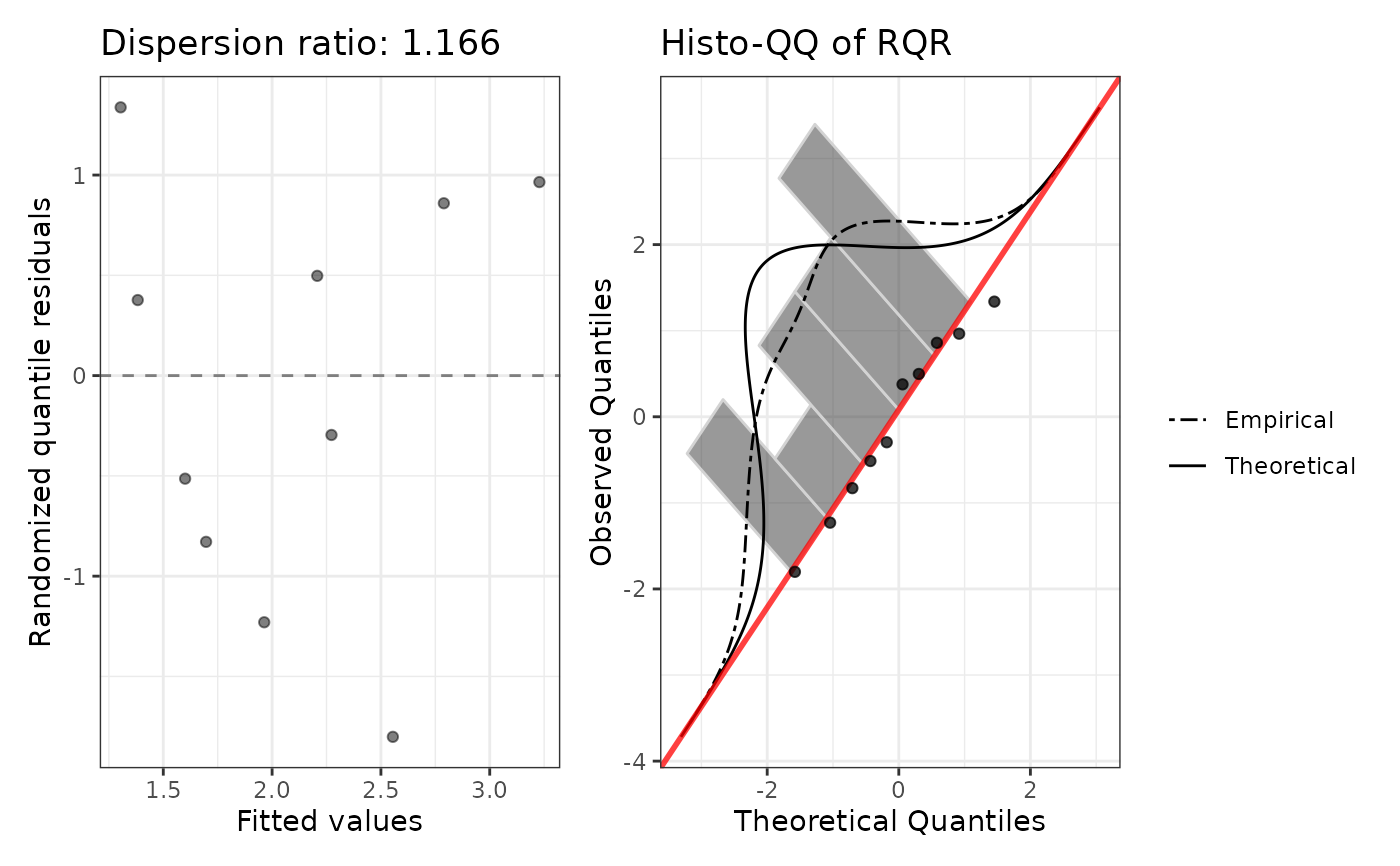
Fits a Poisson GLM and returns model coefficients on the response scale (exponentiated), randomized quantile residuals (RQR), a Pearson dispersion ratio, and a two-panel diagnostic plot.
Arguments
- formula
A model formula (e.g.
y ~ x1 + x2). The response must be a non-negative integer count variable.- data
A data frame containing the variables in
formula.- ...
Additional arguments passed to
stats::glm().
Value
An object of class c("poissonGLM", "countGLMfit"), a list with:
callThe matched call.
modelThe underlying stats::glm fit object.
coefficientsA data frame with columns
term,exp.coef,lower.95,upper.95(all on the response/exponentiated scale).diagnosticsA list with:
rqrNumeric vector of randomized quantile residuals.
dispersion_ratioPearson chi-squared / df.residual. Values substantially above 1 (rule of thumb: > 1.5) suggest overdispersion; consider
negbinGLM().plotA patchwork ggplot: fitted values vs RQR (left) and normal Q-Q of RQR (right).
aicAIC of the fitted model.
Details
Coefficient interpretation: Poisson regression models the log of the expected count. Exponentiating a coefficient gives the multiplicative change in the expected count for a one-unit increase in the predictor, adjusting for simultaneous linear changes in other predictors. For example, 1.5 means a 50% higher expected count.
Condition checking: Inspect diagnostics$dispersion_ratio. A value
near 1 is consistent with the Poisson assumption (mean = variance). The
RQR diagnostic plot should show points scattered randomly around zero with
approximately normal QQ behaviour.
Examples
df <- data.frame(
y = c(0L, 1L, 2L, 3L, 5L, 0L, 2L, 4L, 1L, 3L),
x1 = c(1.2, -0.4, 0.8, -1.1, 2.0, 0.3, -0.9, 1.5, -0.2, 0.7)
)
fit <- poissonGLM(y ~ x1, data = df)
print(fit)
#>
#> Call:
#> poissonGLM(formula = y ~ x1, data = df)
#>
#> Model family: poissonGLM
#>
#> Coefficients (on response scale):
#> term exp.coef lower.95 upper.95
#> (Intercept) 1.7989 1.0683 3.0290
#> x1 1.3396 0.8572 2.0936
#>
#> Dispersion ratio: 1.1658
#> AIC: 39.02
plot(fit)